|
Research Interests
mycobacteria – mycobacterial membrane – membrane transport and signaling – antibiotics – membrane lipids – mycobacterial protein nanocompartments
Disciplines
microbiology – molecular biology – biochemistry – proteomics – organic synthesis – analytical chemistry
Team
- Mariia Zmyslia: mariia.zmyslia(at)ocbc.uni-freiburg.de, Phone 203-6038
- Michael Grimmeisen: michael.grimmeisen(at).ocbc.uni-freiburg.de, Phone 203-6037
- Jürgen Beck: juergen.beck(at).ocbc.uni-freiburg.de, Phone 203-6037
- Yu Zhang: yu.zhang(at)ocbc.uni-freiburg.de, Phone 203-6038
- Melissa Weldle: melissa.weldle(at)pharmazie.uni-freiburg.de, Phone 203-6038
- Trinh Dao: trinh.dao(at)cibss.uni-freiburg.de, Phone 203-6038
- Eleanor Fritz: eleanor.fritz(at)ocbc.uni-freiburg.de, Phone 203-6038
- Julian Boele: julian.boele(at)pharmazie.uni-freiburg.de, Phone 203-6038
- Claudia Jessen-Trefzer: claudia.jessen-trefzer(at)pharmazie.uni-freiburg.de, Phone 203-6074
Projects
 ABC-transporter and Proteomics - Currently we investigate different species of the mycobacterium genus using a lipoproteomics approach. We are especially interested in classifying the mycobacterial transportome in the context of mycobacterial infections. ABC-transporter and Proteomics - Currently we investigate different species of the mycobacterium genus using a lipoproteomics approach. We are especially interested in classifying the mycobacterial transportome in the context of mycobacterial infections.
Li‡, Müller‡, Fröhlich, Gorka, Zhang, Groß, Schilling, Einsle, Jessen-Trefzer*, Cell Chemical Biology, 2019, DOI: 10.1016/j.chembiol.2019.03.002 ‡ equal contribution
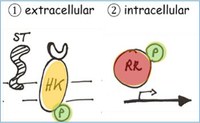 Membrane Signaling via Two - Component Systems - Mycobacteria possess several two-component systems, we are currently studying MSMEG_0244/0246 and its potential interactions with the heme binding protein MSMEG_0243. Membrane Signaling via Two - Component Systems - Mycobacteria possess several two-component systems, we are currently studying MSMEG_0244/0246 and its potential interactions with the heme binding protein MSMEG_0243.
Li, Gašparovič, Weng, Chen, Korduláková* & Jessen-Trefzer*, Frontiers in Microbiology, 2020, DOI: 10.3389/fmicb.2020.570606
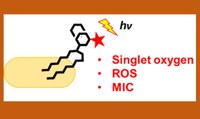 Lipid Anchored Photosensitizers - In order to fight mycobacterial infections at a new level, we develop novel photosensitizers that are actively incorporated into the mycobacterial membrane via the enzyme complex Ag85. Lipid Anchored Photosensitizers - In order to fight mycobacterial infections at a new level, we develop novel photosensitizers that are actively incorporated into the mycobacterial membrane via the enzyme complex Ag85.
Dutta, Chaudahary, Wang, Záhorszka, Forbac, Lohner, Jessen, Argawal, Korduláková, Jessen-Trefzer*, ACS Central Science, 2019, DOI: 10.1021/acscentsci.8b00962
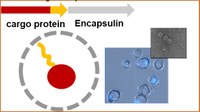 Encapsulins – Protein Nanocompartments in Catalysis - We synthesize organometallic catalysts to covalently incorporate into protein nanocompartments via engineered protein tags. These functional encapsulins are fascinating tools for synthetic biology or pro-drug approaches. Encapsulins – Protein Nanocompartments in Catalysis - We synthesize organometallic catalysts to covalently incorporate into protein nanocompartments via engineered protein tags. These functional encapsulins are fascinating tools for synthetic biology or pro-drug approaches.
Lohner, Zmyslia, Thurn, Pape, Gerasimaite, Keller-Findeisen, Groeer, Deuringer, Süss, Walther, Hell, Lukinavicius, Hugel, Jessen-Trefzer*, Angewandte Chemie, 2021, https://doi.org/10.1002/anie.202110327
Complete List of Publications
- Zmyslia M, Capper MJ, Grimmeisen M, Sartory K, Deuringer B, Abdelsalam M, Shen K, Jung M, Sippl W, Koch HG, Kaul L, Suess R, Koehnke J*, Jessen-Trefzer C*, A nanoengineered tandem nitroreductase: designing a robust prodrug-activating nanoreactor, RSC Chemical Biology, 2024, DOI: 10.1039/d4cb00127c.
- Abdelsalam M, Zmyslia M, Schmidtkunz K, Vecchio A, Hilscher S, Ibrahim HS, Schutkowski M, Jung M, Jessen-Trefzer C*, Sippl W*. Design and synthesis of bioreductive prodrugs of class I histone deacetylase inhibitors and their biological evaluation in virally transfected acute myeloid leukemia cells. Archiv der Pharmazie (Weinheim), 2024, 357(2):e2300536. https://doi: 10.1002/ardp.202300536.
- Zmyslia M, Fröhlich K, Dao T, Schmidt A, Jessen-Trefzer C*, Deep Proteomic Investigation of Metabolic Adaptation in Mycobacteria under Different Growth Conditions, Proteomes, 2023, 11 (4), 39, https://doi: 10.3390/proteomes11040039.
- Grimmeisen M, Jessen-Trefzer C*. Increasing the selectivity of light-active antimicrobial agents - or how to get a photosensitizer to the desired target. ChemBioChem, 2023, 24, 18, e202300177. doi.org/10.1002/cbic.202300177.
- Ebensperger P, Zmyslia M, Lohner P, Braunreuther J, Deuringer B, Becherer A, Süss R, Fischer A, Jessen-Trefzer C*, A bimetal-catalyzed cascade reaction in an engineered protein cage, Angewandte Chemie International Edition, 2023, https://doi.org/10.1002/anie.202218413
- Hacker SM*, Jessen-Trefzer C*. Chemical biology in drug discovery. Biological Chemistry, 2022, 403:361–362.
- Ebensperger P, Jessen-Trefzer C*, Artificial Metalloenzymes in a Nutshell: The Quartet for Efficient Catalysis, Biological Chemistry, 2021, 403:403–412.
-
Lohner P, Zmyslia M, Thurn J, Pape JK, Gerasimaite R, Keller-Findeisen J, Groeer S, Deuringer B, Süss R, Walther A, Lukinavicius G, Hell SW, Hugel T, Jessen-Trefzer C*, Inside a shell - Organometallic catalysis inside encapsulin nano-reactors, Angewandte Chemie International Edition, 2021, https://doi.org/10.1002/anie.202110327 *corresponding author
-
Wang X, Bittner T, Milanov M, Kaul L, Mundinger S, Koch H-G*, Jessen-Trefzer C*, and Jessen HJ*, Pyridinium Modified Anthracenes and Their Endoperoxides Provide a Tunable Scaffold with Activity against Gram-Positive and Gram-Negative Bacteria. ACS Infectious Disease, 2021, 7, 8, 2073–2080. *corresponding author
- Li M, Gašparovič H, Weng X, Chen S, Korduláková J*, Jessen-Trefzer C*: The two-component locus MSMEG_0244/0246 together with MSMEG_0243 affects biofilm assembly in M. smegmatis correlating with changes in phosphatidylinositol mannosides acylation. Frontiers in Microbiology, 2020, doi: 10.3389/fmicb.2020.570606 *corresponding author
- Unzue A, Jessen-Trefzer C, Spiliotopoulos D, Gaudio E, Tarantelli C, Dong J, Zhao H, Leibl J, Zahler S, Bernasconi E, Sartori G, Cascione L, Bertoni F, Sledz P, Caflisch A, Nevado C: Understanding the Mechanism of Action of Pyrrolo[3,2-b]quinoxaline-derivatives as Kinase Inhibitors. RSC Medicinal Chemistry, 2020,11, 665-675.
- Haas TM, Qiu D, Häner M, Angebauer L, Ripp A, Singh J, Koch HG, Jessen-Trefzer C, Jessen HJ: Four phosphates at one blow: access to pentaphosphorylated magic spot nucleotides and their analysis by capillary electrophoresis. Journal of Organic Chemistry, 2020, 85(22):14496-14506.
- Li M, Muller C, Frohlich K, Gorka O, Zhang L, Gross O, Schilling O, Einsle O, Jessen-Trefzer C*: Detection and Characterization of a Mycobacterial L-Arabinofuranose ABC Transporter Identified with a Rapid Lipoproteomics Protocol. Cell Chemical Biology, 2019; 26 (6): 852-862.e6 *corresponding author
- Dutta A, Choudhary E, Wang X, Záhorszka M, Forbak M, Lohner P, Jessen H, Agarwal N, Korduláková J, Jessen-Trefzer C*: Trehalose Conjugation Enhances Toxicity of Photosensitizers against Mycobacteria ACS Central Science, 2019; 5 (4): 644–650. *corresponding author
- Haas TM, Ebensperger P, Eisenbeis VB, Nopper C, Durr T, Jork N, Steck N, Jessen-Trefzer C, Jessen HJ: Magic spot nucleotides: tunable target-specific chemoenzymatic synthesis. Chemical Communication, 2019; 55, 5339-5342.
- Zhang S, Zhu J, Zechel DL, Jessen-Trefzer C, Eastman RT, Paululat T, Bechthold A: New WS9326A Derivatives and One New Annimycin Derivative with Antimalarial Activity are Produced by Streptomyces asterosporus DSM 41452 and Its Mutant. ChemBioChem, 2018; 19 (3): 272-279.
- Greule A, Marolt M, Deubel D, Peintner I, Zhang S, Jessen-Trefzer C, De Ford C, Burschel S, Li SM, Friedrich T, Merfort I, Ludeke S, Bisel P, Muller M, Paululat T, Bechthold A: Wide Distribution of Foxicin Biosynthetic Gene Clusters in Streptomyces Strains An Unusual Secondary Metabolite with Various Properties. Frontiers in Microbiology, 2017; 8: 221-221.
Postdoc and Doctoral Studies (Birth name: Trefzer)
- Herdy B, Karonitsch T, Vladimer GI, Tan CS, Stukalov A, Trefzer C, Bigenzahn JW, Theil T, Holinka J, Kiener HP, Colinge J, Bennett KL, Superti-Furga G: The RNA-binding protein HuR/ELAVL1 regulates IFN-beta mRNA abundance and the type I IFN response. European Journal of Immunology, 2015; 45 (5): 1500-1511.
- Hofer A, Cremosnik GS, Muller AC, Giambruno R, Trefzer C, Superti-Furga G, Bennett KL, Jessen HJ: A Modular Synthesis of Modified Phosphoanhydrides. Chemical European Journal, 2015; 21 (28): 10116-10122.
- Licciardello MP, Müllner MK, Dürnberger G, Kerzendorfer C, Boidol B, Trefzer C, Sdelci S, Berg T, Penz T, Schuster M, Bock C, Kralovics R, Superti-Furga G, Colinge J, Nijman SM, Kubicek S. NOTCH1 activation in breast cancer confers sensitivity to inhibition of SUMOylation. Oncogene, 2015; 34 (29) :3780-90.
- Muellner MK, Mair B, Ibrahim Y, Kerzendorfer C, Lechtermann H, Trefzer C, Klepsch F, Muller AC, Leitner E, Macho-Maschler S, Superti-Furga G, Bennett KL, Baselga J, Rix U, Kubicek S, Colinge J, Serra V, Nijman SM: Targeting a cell state common to triple-negative breast cancers. Molecular Systems Biology, 2015; 11 (1): 789-789.
- Winter GE, Radic B, Mayor-Ruiz C, Blomen VA, Trefzer C, Kandasamy RK, Huber KVM, Gridling M, Chen D, Klampfl T, Kralovics R, Kubicek S, Fernandez-Capetillo O, Brummelkamp TR, Superti-Furga G: The solute carrier SLC35F2 enables YM155-mediated DNA damage toxicity. Nature Chemical Biology, 2014; 10 (9): 768-773.
- Wang F, Sambandan D, Halder R, Wang J, Batt SM, Weinrick B, Ahmad I, Yang P, Zhang Y, Kim J, Hassani M, Huszar S, Trefzer C, Ma Z, Kaneko T, Mdluli KE, Franzblau S, Chatterjee AK, Johnsson K, Mikusova K, Besra GS, Futterer K, Robbins SH, Barnes SW, Walker JR, Jacobs WR Jr, Schultz PG: Identification of a small molecule with activity against drug-resistant and persistent tuberculosis. PNAS, 2013; 110 (27): E2510-E2517.
- Trefzer C, Skovierova H, Buroni S, Bobovska A, Nenci S, Molteni E, Pojer F, Pasca MR, Makarov V, Cole ST, Riccardi G, Mikusova K, Johnsson K: Benzothiazinones are suicide inhibitors of mycobacterial decaprenylphosphoryl-beta-D-ribofuranose 2’-oxidase DprE1. JACS, 2012; 134 (2): 912-915.
- Trefzer C, Rengifo-Gonzalez M, Hinner MJ, Schneider P, Makarov V, Cole ST, Johnsson K: Benzothiazinones: prodrugs that covalently modify the decaprenylphosphoryl-beta-D-ribose 2’-epimerase DprE1 of Mycobacterium tuberculosis. JACS, 2010; 132 (39): 13663-13665.
or FreiDok
|



 ABC-transporter and Proteomics - Currently we investigate different species of the mycobacterium genus using a lipoproteomics approach. We are especially interested in classifying the mycobacterial transportome in the context of mycobacterial infections.
ABC-transporter and Proteomics - Currently we investigate different species of the mycobacterium genus using a lipoproteomics approach. We are especially interested in classifying the mycobacterial transportome in the context of mycobacterial infections. Membrane Signaling via Two - Component Systems - Mycobacteria possess several two-component systems, we are currently studying MSMEG_0244/0246 and its potential interactions with the heme binding protein MSMEG_0243.
Membrane Signaling via Two - Component Systems - Mycobacteria possess several two-component systems, we are currently studying MSMEG_0244/0246 and its potential interactions with the heme binding protein MSMEG_0243. Lipid Anchored Photosensitizers - In order to fight mycobacterial infections at a new level, we develop novel photosensitizers that are actively incorporated into the mycobacterial membrane via the enzyme complex Ag85.
Lipid Anchored Photosensitizers - In order to fight mycobacterial infections at a new level, we develop novel photosensitizers that are actively incorporated into the mycobacterial membrane via the enzyme complex Ag85. Encapsulins – Protein Nanocompartments in Catalysis - We synthesize organometallic catalysts to covalently incorporate into protein nanocompartments via engineered protein tags. These functional encapsulins are fascinating tools for synthetic biology or pro-drug approaches.
Encapsulins – Protein Nanocompartments in Catalysis - We synthesize organometallic catalysts to covalently incorporate into protein nanocompartments via engineered protein tags. These functional encapsulins are fascinating tools for synthetic biology or pro-drug approaches.